Chemical Engineering with DNA

Contact: Prof. Robert Grass
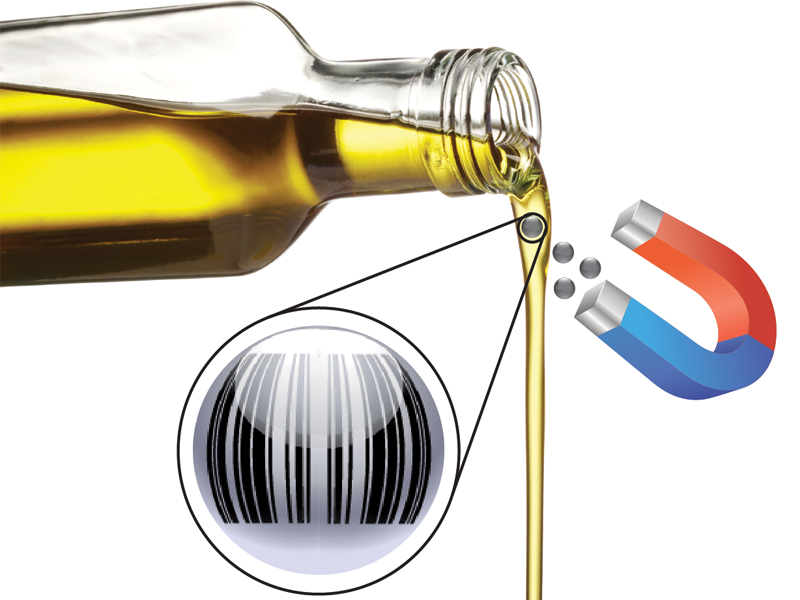
Anti-counterfeit of materials and products can be seen as a specific tracer application, where the information-carrying tracer (or taggant) is incorporated into (or onto) the valuable good. Particle-encapsulated DNA is a promising candidate as an anti-counterfeit tag, as large amounts of information can be encoded, data encryption methods exist for DNA, and the applied materials (glass and DNA) are safe to use and already approved as additives in packaging, food, and pharmaceuticals. Due to the almost unlimited amount of different codes that can be generated and the scalability of the technology, this approach would allow the creation of a global materials labeling platform, where the fate of every raw material and final product can be evaluated. Proof of concept data is already available for the tracing of polymers, foodstuff (milk, cheese, olive oil) [1] and industrial chemical feedstocks [2].
[1] J. Agric. Food Chem., 2015. external page Link
[2] ACS Nano, 2014. external page Link, ETH News article

Contact: Prof. Robert Grass
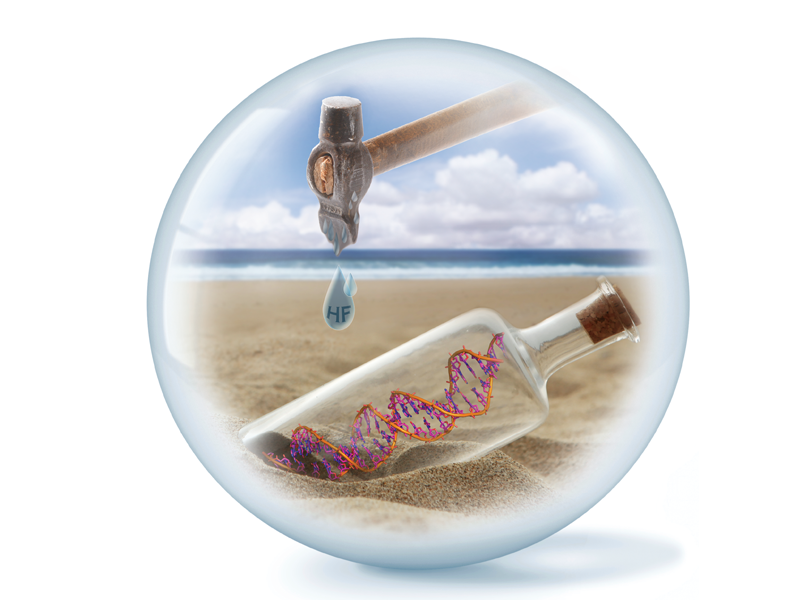
Ancient DNA is preserved in fossils for over a hundred thousand of years. This natural way of DNA preservation inspired us to encapsulate DNA in glass particles (synthetic fossils). Within the particle matrix, the DNA is hermetically sealed and protected against damaging environmental attack, such as reactive oxygen species and high temperatures.[1,2] With an additional layer of TiO2 the encapsulated nucleic acids can be shielded from aggressive UV-irradiation.[3] The recovery of DNA is achieved by dissolving the particles with a diluted fluoride buffer. This technique can be used for short artificial DNA strands, but also for plasmids, genomic DNA[4] and even RNA.[5]
[1] R. N. Grass, W. J. Stark, WO2013143014
[2] Angew. Chem. Int. Ed., 2013. external page Link
[3] J. Mater. Chem. B, 2014. external page Link
[4] Nat. Protoc., 2013. external page Link
[5] Adv. Healthc. Mater., 2015. external page Link

Contact: Prof. Robert Grass
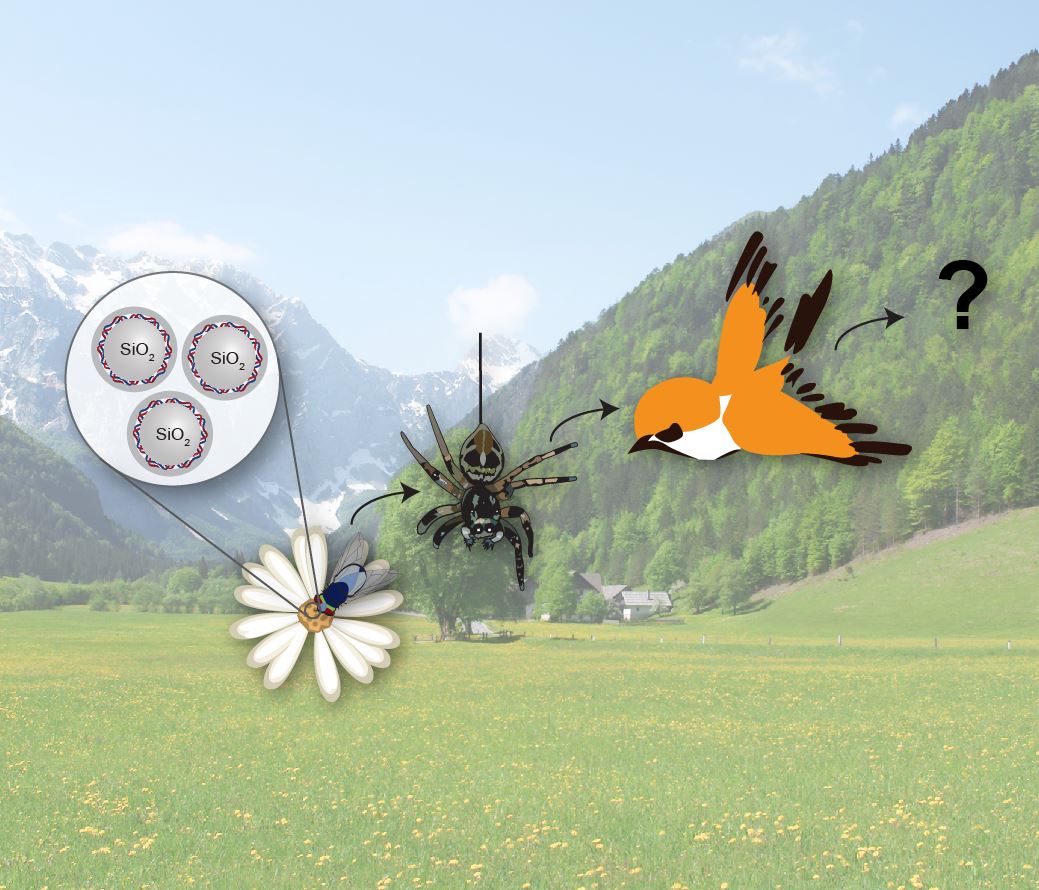
The synthetic fossil particles can be detected and quantified via the encapsulated DNA tag using quantitative real-time polymerase chain reaction (qPCR), allowing the application of these particles as tracer tools. Due to the excellent specificity and sensitivity of qPCR, particles can be detected down to ppb concentration levels (= µg/kg) and by the use of digital PCR even individual nanoparticles can be counted one by one. [6] This technology can be used to investigate the fate of silica nanoparticles in living cells (nanotoxicology) [7] and the environment (e.g. wastewater treatment).
Additionally, the particles can be used to follow (trace) the flow and distribution of liquids (river flow and subterranean reservoir characterization [8]), chemicals such as pesticide spread in the environment [9] and even the ecological feeding relationships of animals. [10] Current work in this field includes the development of faster and more mobile DNA quantification systems.
[6] ACS Nano, 2015. external page Link, external page Video
[7] Chem. Commun., 2014. external page Link
[8] Environ. Sci. Technol., 2018. external page Link
[9] Environ. Sci. Technol. Lett., 2016. external page Link
[10] Mol. Ecol. Resourc., 2015. external page Link
Contact: Prof. Robert Grass
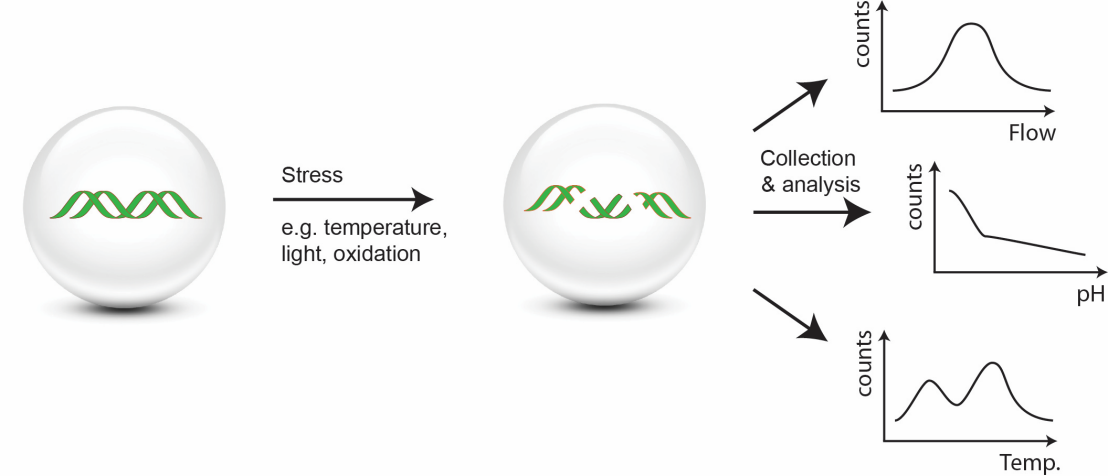
The ability of DNA-based tracers to store information makes them attractive for performing distributed measurements and delivering localized information upon recollection. To achieve this, DNA is modified so that it is vulnerable to a specific external stimulus. After exposure, the particles are recollected and the change in DNA composition is related to an external stimulus [11]. We have demonstrated that smart DNA-based tracers can measure temperature [12], oxidative stress, and light intensity or duration [13] in challenging environments (e.g. single cells).
[11] G. Mikutis, M. Puddu, R. N. Grass, W. J. Stark, EP15164214
[12] Small, 2016. external page Link
[13] Adv. Mater., 2016. external page Link

Contact: Prof. Robert Grass
Some of the oldest information we currently have access to is the genomic information of ancient species stored as DNA in bone fossils (e.g. the 700’000 year old genome of a horse). Since DNA, encapsulated in our synthetic fossils, is as stable as DNA in natural fossils, we have investigated the use of encapsulated DNA for long-term digital information storage. As a prototype we stored Archimedes’ Methods of Mechanical Theorems and the Swiss Federal Charter (total of 83 kB) as synthetically synthesized DNA [14]. By combining DNA encapsulation in artificial fossils and forward error correction coding digital data could be recovered error-free after 2000 years of simulated ambient-temperature storage. Current work in this area includes engineering methods to decrease the cost of array-based DNA synthesis, which is required to make DNA data storage competitive with current data storage methods (e.g. magnetic storage) [15, 16].
[14] Angew. Chem. Int. Ed., 2015. external page Link
[15] Adv. Funct. Mater., 2019. external page Link
[16] Sci. Reports, 2019. external page Link